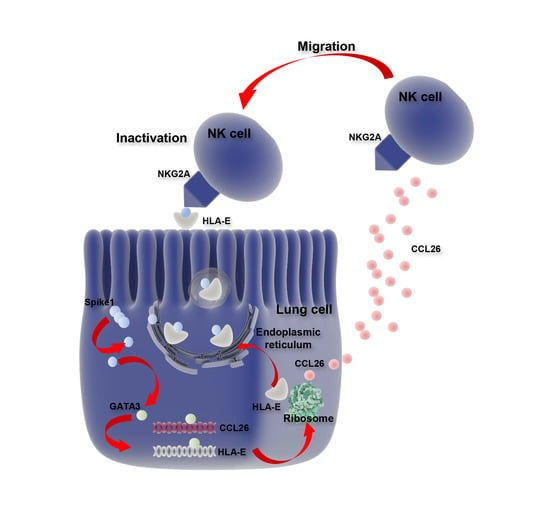
SARS-CoV-2 Spike 1 Protein Controls Natural Killer Cell Activation via the HLA-E/NKG2A Pathway
Abstract
:1. Introduction
2. Materials and Methods
2.1. NK Cells Purification
2.2. Viral RNA Detection
2.3. Cell Culture
2.4. Flow Cytometry
2.5. Immunofluorescence Assay
2.6. Cell Migration Assay
2.7. Protein Transfection
2.8. Lactate Dehydrogenase (LDH) Assay
2.9. Degranulation Analysis
2.10. Carboxyfluorescein Diacetate Succinimidyl Ester (CFSE) Analysis
2.11. IFN-gamma ELISA Assay
2.12. Real-Time PCR
2.13. HLA-E Binding Prediction
2.14. Statistical Analysis
2.15. Data Availability
3. Results
3.1. Spike Proteins Induce NK Cell Migration
3.2. Spike Proteins Did Not Modify NK Cell Cytotoxicity
3.3. SARS-CoV-2 Spike 1 Protein Modifies NK Cell Cytotoxicity When Presented by Lung Epithelial Cells
3.4. Spike 1 Peptide Is Presented by Lung Epithelial Cells via HLA-E Molecules
3.5. HLA-E/NKG2A/CD94 Interaction Is Responsible for NK Cells’ Exhaustion
4. Discussion
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Zhu, N.; Zhang, D.; Wang, W.; Li, X.; Yang, B.; Song, J.; Zhao, X.; Huang, B.; Shi, W.; Lu, R.; et al. A Novel Coronavirus from Patients with Pneumonia in China, 2019. N. Engl. J. Med. 2020, 382, 727–733. [Google Scholar] [CrossRef] [PubMed]
- Li, Q.; Guan, X.; Wu, P.; Wang, X.; Zhou, L.; Tong, Y.; Ren, R.; Leung, K.S.M.; Lau, E.H.Y.; Wong, J.Y.; et al. Early Transmission Dynamics in Wuhan, China, of Novel Coronavirus-Infected Pneumonia. N. Engl. J. Med. 2020, 382, 1199–1207. [Google Scholar] [CrossRef] [PubMed]
- Zhou, P.; Yang, X.L.; Wang, X.G.; Hu, B.; Zhang, L.; Zhang, W.; Si, H.R.; Zhu, Y.; Li, B.; Huang, C.L.; et al. A pneumonia outbreak associated with a new coronavirus of probable bat origin. Nature 2020, 579, 270–273. [Google Scholar] [CrossRef] [Green Version]
- Guan, W.J.; Ni, Z.Y.; Hu, Y.; Liang, W.H.; Ou, C.Q.; He, J.X.; Liu, L.; Shan, H.; Lei, C.L.; Hui, D.S.C.; et al. Clinical Characteristics of Coronavirus Disease 2019 in China. N. Engl. J. Med. 2020, 382, 1708–1720. [Google Scholar] [CrossRef] [PubMed]
- Qin, C.; Zhou, L.; Hu, Z.; Zhang, S.; Yang, S.; Tao, Y.; Xie, C.; Ma, K.; Shang, K.; Wang, W.; et al. Dysregulation of immune response in patients with COVID-19 in Wuhan, China. Clin. Infect. Dis. 2020. [Google Scholar] [CrossRef]
- Pedersen, S.F.; Ho, Y.C. SARS-CoV-2: A storm is raging. J. Clin. Investig. 2020, 130, 2202–2205. [Google Scholar] [CrossRef] [PubMed]
- Gornalusse, G.G.; Hirata, R.K.; Funk, S.E.; Riolobos, L.; Lopes, V.S.; Manske, G.; Prunkard, D.; Colunga, A.G.; Hanafi, L.A.; Clegg, D.O.; et al. HLA-E-expressing pluripotent stem cells escape allogeneic responses and lysis by NK cells. Nat. Biotechnol. 2017, 35, 765–772. [Google Scholar] [CrossRef] [Green Version]
- Tan, L.; Wang, Q.; Zhang, D.; Ding, J.; Huang, Q.; Tang, Y.Q.; Wang, Q.; Miao, H. Lymphopenia predicts disease severity of COVID-19: A descriptive and predictive study. Signal. Transduct. Target. Ther. 2020, 5, 33. [Google Scholar] [CrossRef]
- Vivier, E.; Tomasello, E.; Baratin, M.; Walzer, T.; Ugolini, S. Functions of natural killer cells. Nat. Immunol. 2008, 9, 503–510. [Google Scholar] [CrossRef]
- Long, E.O.; Kim, H.S.; Liu, D.; Peterson, M.E.; Rajagopalan, S. Controlling natural killer cell responses: Integration of signals for activation and inhibition. Annu. Rev. Immunol. 2013, 31, 227–258. [Google Scholar] [CrossRef] [Green Version]
- Houchins, J.P.; Lanier, L.L.; Niemi, E.C.; Phillips, J.H.; Ryan, J.C. Natural killer cell cytolytic activity is inhibited by NKG2-A and activated by NKG2-C. J. Immunol. 1997, 158, 3603–3609. [Google Scholar] [PubMed]
- Nguyen, S.; Beziat, V.; Dhedin, N.; Kuentz, M.; Vernant, J.P.; Debre, P.; Vieillard, V. HLA-E upregulation on IFN-gamma-activated AML blasts impairs CD94/NKG2A-dependent NK cytolysis after haplo-mismatched hematopoietic SCT. Bone Marrow Transplant. 2009, 43, 693–699. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Miller, J.D.; Weber, D.A.; Ibegbu, C.; Pohl, J.; Altman, J.D.; Jensen, P.E. Analysis of HLA-E peptide-binding specificity and contact residues in bound peptide required for recognition by CD94/NKG2. J. Immunol. 2003, 171, 1369–1375. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Braud, V.; Jones, E.Y.; McMichael, A. The human major histocompatibility complex class Ib molecule HLA-E binds signal sequence-derived peptides with primary anchor residues at positions 2 and 9. Eur. J. Immunol. 1997, 27, 1164–1169. [Google Scholar] [CrossRef] [PubMed]
- Lee, N.; Llano, M.; Carretero, M.; Ishitani, A.; Navarro, F.; Lopez-Botet, M.; Geraghty, D.E. HLA-E is a major ligand for the natural killer inhibitory receptor CD94/NKG2A. Proc. Natl. Acad. Sci. USA 1998, 95, 5199–5204. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lazetic, S.; Chang, C.; Houchins, J.P.; Lanier, L.L.; Phillips, J.H. Human natural killer cell receptors involved in MHC class I recognition are disulfide-linked heterodimers of CD94 and NKG2 subunits. J. Immunol. 1996, 157, 4741–4745. [Google Scholar]
- Marci, R.; Gentili, V.; Bortolotti, D.; Lo Monte, G.; Caselli, E.; Bolzani, S.; Rotola, A.; Di Luca, D.; Rizzo, R. Presence of HHV-6A in Endometrial Epithelial Cells from Women with Primary Unexplained Infertility. PLoS ONE 2016, 11, e0158304. [Google Scholar] [CrossRef]
- Mirjacic Martinovic, K.; Babovic, N.; Dzodic, R.; Jurisic, V.; Matkovic, S.; Konjevic, G. Favorable in vitro effects of combined IL-12 and IL-18 treatment on NK cell cytotoxicity and CD25 receptor expression in metastatic melanoma patients. J. Transl. Med. 2015, 13, 120. [Google Scholar] [CrossRef] [Green Version]
- Nomura, S.; Takahashi, H.; Suzuki, J.; Kuwahara, M.; Yamashita, M.; Sawasaki, T. Pyrrothiogatain acts as an inhibitor of GATA family proteins and inhibits Th2 cell differentiation in vitro. Sci. Rep. 2019, 9, 17335. [Google Scholar] [CrossRef] [Green Version]
- Bortolotti, D.; Gentili, V.; Rotola, A.; Cultrera, R.; Marci, R.; Di Luca, D.; Rizzo, R. HHV-6A infection of endometrial epithelial cells affects immune profile and trophoblast invasion. Am. J. Reprod. Immunol. 2019, 82, e13174. [Google Scholar] [CrossRef] [Green Version]
- Bortolotti, D.; Gentili, V.; Caselli, E.; Sicolo, M.; Soffritti, I.; D’Accolti, M.; Barao, I.; Rotola, A.; Di Luca, D.; Rizzo, R. DNA Sensors’ Signaling in NK Cells During HHV-6A, HHV-6B and HHV-7 Infection. Front. Microbiol. 2020, 11, 226. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Inngjerdingen, M.; Rolstad, B.; Ryan, J.C. Activating and inhibitory Ly49 receptors modulate NK cell chemotaxis to CXC chemokine ligand (CXCL) 10 and CXCL12. J. Immunol. 2003, 171, 2889–2895. [Google Scholar] [CrossRef] [Green Version]
- Rizzo, R.; Gentili, V.; Casetta, I.; Caselli, E.; De Gennaro, R.; Granieri, E.; Cassai, E.; Di Luca, D.; Rotola, A. Altered natural killer cells’ response to herpes virus infection in multiple sclerosis involves KIR2DL2 expression. J. Neuroimmunol. 2012, 251, 55–64. [Google Scholar] [CrossRef]
- Zitvogel, L. Dendritic and natural killer cells cooperate in the control/switch of innate immunity. J. Exp. Med. 2002, 195, F9–F14. [Google Scholar] [CrossRef] [PubMed]
- Mwimanzi, P.; Markle, T.J.; Ueno, T.; Brockman, M.A. Human leukocyte antigen (HLA) class I down-regulation by human immunodeficiency virus type 1 negative factor (HIV-1 Nef): What might we learn from natural sequence variants? Viruses 2012, 4, 1711–1730. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Stevens, J.; Joly, E.; Trowsdale, J.; Butcher, G.W. Peptide binding characteristics of the non-classical class Ib MHC molecule HLA-E assessed by a recombinant random peptide approach. BMC Immunol. 2001, 2, 5. [Google Scholar] [CrossRef]
- Caramori, G.; Lim, S.; Ito, K.; Tomita, K.; Oates, T.; Jazrawi, E.; Chung, K.F.; Barnes, P.J.; Adcock, I.M. Expression of GATA family of transcription factors in T-cells, monocytes and bronchial biopsies. Eur. Respir. J. 2001, 18, 466–473. [Google Scholar] [CrossRef] [Green Version]
- Zheng, W.; Flavell, R.A. The transcription factor GATA-3 is necessary and sufficient for Th2 cytokine gene expression in CD4 T cells. Cell 1997, 89, 587–596. [Google Scholar] [CrossRef] [Green Version]
- Sun, Q.; Yang, X.; Zhong, B.; Jiao, F.; Li, C.; Li, D.; Lan, X.; Sun, J.; Lu, S. Upregulated protein arginine methyltransferase 1 by IL-4 increases eotaxin-1 expression in airway epithelial cells and participates in antigen-induced pulmonary inflammation in rats. J. Immunol. 2012, 188, 3506–3512. [Google Scholar] [CrossRef] [Green Version]
- Zheng, M.; Gao, Y.; Wang, G.; Song, G.; Liu, S.; Sun, D.; Xu, Y.; Tian, Z. Functional exhaustion of antiviral lymphocytes in COVID-19 patients. Cell Mol. Immunol. 2020, 17, 533–535. [Google Scholar] [CrossRef] [Green Version]
- Antonioli, L.; Fornai, M.; Pellegrini, C.; Blandizzi, C. NKG2A and COVID-19: Another brick in the wall. Cell Mol. Immunol. 2020. [Google Scholar] [CrossRef] [PubMed]
- Yaqinuddin, A.; Kashir, J. Innate immunity in COVID-19 patients mediated by NKG2A receptors, and potential treatment using Monalizumab, Cholroquine, and antiviral agents. Med. Hypotheses 2020, 140, 109777. [Google Scholar] [CrossRef] [PubMed]
- Jewett, A. The Potential Effect of Novel Coronavirus SARS-CoV-2 on NK Cells; A Perspective on Potential Therapeutic Interventions. Front. Immunol. 2020, 11, 1692. [Google Scholar] [CrossRef] [PubMed]
- Market, M.A.L.; Martel, A.B.; Bastin, D.; Olanubi, O.; Tennakoon, G.; Boucher, D.M.; Ng, J.; Ardolino, M.; Auer, R.C. Flattening the COVID-19 Curve With Natural Killer Cell Based Immunotherapies. Front. Immunol. 2020, 11. [Google Scholar] [CrossRef] [PubMed]
- Hoffmann, M.; Kleine-Weber, H.; Schroeder, S.; Kruger, N.; Herrler, T.; Erichsen, S.; Schiergens, T.S.; Herrler, G.; Wu, N.H.; Nitsche, A.; et al. SARS-CoV-2 Cell Entry Depends on ACE2 and TMPRSS2 and Is Blocked by a Clinically Proven Protease Inhibitor. Cell 2020, 181, 271–280.e8. [Google Scholar] [CrossRef] [PubMed]
- de Lang, A.; Osterhaus, A.D.; Haagmans, B.L. Interferon-gamma and interleukin-4 downregulate expression of the SARS coronavirus receptor ACE2 in Vero E6 cells. Virology 2006, 353, 474–481. [Google Scholar] [CrossRef] [Green Version]
- Frieman, M.; Heise, M.; Baric, R. SARS coronavirus and innate immunity. Virus Res. 2008, 133, 101–112. [Google Scholar] [CrossRef]
- Anfossi, N.; Andre, P.; Guia, S.; Falk, C.S.; Roetynck, S.; Stewart, C.A.; Breso, V.; Frassati, C.; Reviron, D.; Middleton, D.; et al. Human NK cell education by inhibitory receptors for MHC class I. Immunity 2006, 25, 331–342. [Google Scholar] [CrossRef]
- Grote, D.; Boualia, S.K.; Souabni, A.; Merkel, C.; Chi, X.; Costantini, F.; Carroll, T.; Bouchard, M. Gata3 acts downstream of beta-catenin signaling to prevent ectopic metanephric kidney induction. PLoS Genet. 2008, 4, e1000316. [Google Scholar] [CrossRef] [Green Version]
- El-Shazly, A.E.; Doloriert, H.C.; Bisig, B.; Lefebvre, P.P.; Delvenne, P.; Jacobs, N. Novel cooperation between CX3CL1 and CCL26 inducing NK cell chemotaxis via CX3CR1: A possible mechanism for NK cell infiltration of the allergic nasal tissue. Clin. Exp. Allergy 2013, 43, 322–331. [Google Scholar] [CrossRef]
- Hirano, N.; Butler, M.O.; Xia, Z.; Ansen, S.; von Bergwelt-Baildon, M.S.; Neuberg, D.; Freeman, G.J.; Nadler, L.M. Engagement of CD83 ligand induces prolonged expansion of CD8+ T cells and preferential enrichment for antigen specificity. Blood 2006, 107, 1528–1536. [Google Scholar] [CrossRef] [PubMed]
- Ulianich, L.; Terrazzano, G.; Annunziatella, M.; Ruggiero, G.; Beguinot, F.; Di Jeso, B. ER stress impairs MHC Class I surface expression and increases susceptibility of thyroid cells to NK-mediated cytotoxicity. Biochim. Biophys. Acta 2011, 1812, 431–438. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- van Hall, T.; Andre, P.; Horowitz, A.; Ruan, D.F.; Borst, L.; Zerbib, R.; Narni-Mancinelli, E.; van der Burg, S.H.; Vivier, E. Monalizumab: Inhibiting the novel immune checkpoint NKG2A. J. Immunother. Cancer 2019, 7, 263. [Google Scholar] [CrossRef] [PubMed]
- Masselli, E.V.M.; Carubbi, C.; Pozzi, G.; Presta, V.; Mirandola, P.; Vitale, M. NK cells: A double edge sword against SARS-CoV-2. Adv. Biol. Regul. 2020, 77. [Google Scholar] [CrossRef] [PubMed]







© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Bortolotti, D.; Gentili, V.; Rizzo, S.; Rotola, A.; Rizzo, R. SARS-CoV-2 Spike 1 Protein Controls Natural Killer Cell Activation via the HLA-E/NKG2A Pathway. Cells 2020, 9, 1975. https://doi.org/10.3390/cells9091975
Bortolotti D, Gentili V, Rizzo S, Rotola A, Rizzo R. SARS-CoV-2 Spike 1 Protein Controls Natural Killer Cell Activation via the HLA-E/NKG2A Pathway. Cells. 2020; 9(9):1975. https://doi.org/10.3390/cells9091975
Chicago/Turabian StyleBortolotti, Daria, Valentina Gentili, Sabrina Rizzo, Antonella Rotola, and Roberta Rizzo. 2020. "SARS-CoV-2 Spike 1 Protein Controls Natural Killer Cell Activation via the HLA-E/NKG2A Pathway" Cells 9, no. 9: 1975. https://doi.org/10.3390/cells9091975