Abstract
Familial Mediterranean fever (FMF) is an autoinflammatory disease caused by homozygous or compound heterozygous gain-of-function mutations in MEFV, which encodes pyrin, an inflammasome protein. Heterozygous carrier frequencies for multiple MEFV mutations are high in several Mediterranean populations, suggesting that they confer selective advantage. Among 2,313 Turkish people, we found extended haplotype homozygosity flanking FMF-associated mutations, indicating evolutionarily recent positive selection of FMF-associated mutations. Two pathogenic pyrin variants independently arose >1,800 years ago. Mutant pyrin interacts less avidly with Yersinia pestis virulence factor YopM than with wild-type human pyrin, thereby attenuating YopM-induced interleukin (IL)-1β suppression. Relative to healthy controls, leukocytes from patients with FMF harboring homozygous or compound heterozygous mutations and from asymptomatic heterozygous carriers released heightened IL-1β specifically in response to Y. pestis. Y. pestis-infected MefvM680I/M680I FMF knock-in mice exhibited IL-1-dependent increased survival relative to wild-type knock-in mice. Thus, FMF mutations that were positively selected in Mediterranean populations confer heightened resistance to Y. pestis.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
Data availability
The datasets generated and/or analyzed during the current study are available from the Supplementary Information and Source Data files in the online version of the paper and also available from the corresponding author on request.
Code availability
Computer code for the forward-time simulations is available at https://github.com/dshriner/forward-time-simulators under the GNU General Public License v.3.0.
References
International FMF Consortium. Ancient missense mutations in a new member of the RoRet gene family are likely to cause familial Mediterranean fever. Cell 90, 797–807 (1997).
French_FMF_Consortium. A candidate gene for familial Mediterranean fever. Nat. Genet. 17, 25–31 (1997).
Chae, J. J. et al. Gain-of-function pyrin mutations induce NLRP3 protein-independent interleukin-1β activation and severe autoinflammation in mice. Immunity 34, 755–768 (2011).
Sohar, E., Gafni, J., Pras, M. & Heller, H. Familial Mediterranean fever. A survey of 470 cases and review of the literature. Am. J. Med. 43, 227–253 (1967).
Touitou, I. The spectrum of Familial Mediterranean Fever (FMF) mutations. Eur. J. of Hum. Genet. 9, 473–483 (2001).
Kirino, Y. et al. Targeted resequencing implicates the familial Mediterranean fever gene MEFV and the Toll-like receptor 4 gene TLR4 in Behçet disease. Proc. Natl Acad. Sci. USA 110, 8134–8139 (2013).
Papadopoulos, V. P., Giaglis, S., Mitroulis, I. & Ritis, K. The population genetics of familial Mediterranean fever: a meta-analysis study. Ann. Hum. Genet. 72, 752–761 (2008).
Yilmaz, E. et al. Mutation frequency of familial Mediterranean fever and evidence for a high carrier rate in the Turkish population. Eur. J. Hum. Genet. 9, 553–555 (2001).
Koshy, R., Sivadas, A. & Scaria, V. Genetic epidemiology of familial Mediterranean fever through integrative analysis of whole genome and exome sequences from Middle East and North Africa. Clin. Genet. 93, 92–102 (2018).
Zvereff, V. V., Faruki, H., Edwards, M. & Friedman, K. J. Cystic fibrosis carrier screening in a North American population. Genet. Med. 16, 539–546 (2014).
Xu, H. et al. Innate immune sensing of bacterial modifications of Rho GTPases by the pyrin inflammasome. Nature 513, 237–241 (2014).
Park, Y. H., Wood, G., Kastner, D. L. & Chae, J. J. Pyrin inflammasome activation and RhoA signaling in the autoinflammatory diseases FMF and HIDS. Nat. Immunol. 17, 914–921 (2016).
Chung, L. K. et al. The Yersinia virulence factor YopM hijacks host kinases to inhibit type III effector-triggered activation of the pyrin inflammasome. Cell Host Microbe 20, 296–306 (2016).
Ratner, D. et al. The Yersinia pestis effector YopM inhibits pyrin inflammasome activation. PLoS Pathog. 12, e1006035 (2016).
Voight, B. F., Kudaravalli, S., Wen, X. & Pritchard, J. K. A map of recent positive selection in the human genome. PLoS Biol. 4, e72 (2006).
Ferrer-Admetlla, A., Liang, M., Korneliussen, T. & Nielsen, R. On detecting incomplete soft or hard selective sweeps using haplotype structure. Mol. Biol. Evol. 31, 1275–1291 (2014).
Gandolfo, L. C., Bahlo, M. & Speed, T. P. Dating rare mutations from small samples with dense marker data. Genetics 197, 1315–1327 (2014).
Moorjani, P. et al. A genetic method for dating ancient genomes provides a direct estimate of human generation interval in the last 45,000 years. Proc. Natl Acad. Sci. USA 113, 5652–5657 (2016).
Chen, H., Hey, J. & Slatkin, M. A hidden Markov model for investigating recent positive selection through haplotype structure. Theor. Popul. Biol. 99, 18–30 (2015).
Hashemi Shahraki, A., Carniel, E. & Mostafavi, E. Plague in Iran: its history and current status. Epidemiol. Health 38, e2016033 (2016).
McDonald, C., Vacratsis, P. O., Bliska, J. B. & Dixon, J. E. The Yersinia virulence factor YopM forms a novel protein complex with two cellular kinases. J. Biol. Chem. 278, 18514–18523 (2003).
Hentschke, M. et al. Yersinia virulence factor YopM induces sustained RSK activation by interfering with dephosphorylation. PLoS ONE 5, e13165 (2010).
Evdokimov, A. G., Anderson, D. E., Routzahn, K. M. & Waugh, D. S. Unusual molecular architecture of the Yersinia pestis cytotoxin YopM: a leucine-rich repeat protein with the shortest repeating unit. J. Mol. Biol. 312, 807–821 (2001).
McPhee, J. B., Mena, P. & Bliska, J. B. Delineation of regions of the Yersinia YopM protein required for interaction with the RSK1 and PRK2 host kinases and their requirement for interleukin-10 production and virulence. Infect. Immun. 78, 3529–3539 (2010).
McCoy, M. W., Marre, M. L., Lesser, C. F. & Mecsas, J. The C-terminal tail of Yersinia pseudotuberculosis YopM is critical for interacting with RSK1 and for virulence. Infect. Immun. 78, 2584–2598 (2010).
Aubert, D. F. et al. A Burkholderia type VI effector deamidates Rho GTPases to activate the pyrin inflammasome and trigger inflammation. Cell Host Microbe 19, 664–674 (2016).
Allison, A. C. Protection afforded by sickle-cell trait against subtertian malarial infection. Br. Med. J. 1, 290–294 (1954).
Chae, J. J. et al. Isolation, genomic organization, and expression analysis of the mouse and rat homologs of MEFV, the gene for familial Mediterranean fever. Mamm. Genome 11, 428–435 (2000).
Osei-Owusu, P., Charlton, T. M., Kim, H. K., Missiakas, D. & Schneewind, O. FPR1 is the plague receptor on host immune cells. Nature 574, 57–62 (2019).
Achtman, M. et al. Yersinia pestis, the cause of plague, is a recently emerged clone of Yersinia pseudotuberculosis. Proc. Natl Acad. Sci. USA 96, 14043–14048 (1999).
Wagner, D. M. et al. Yersinia pestis and the plague of Justinian 541–543 AD: a genomic analysis. Lancet Infect. Dis. 14, 319–326 (2014).
Achtman, M. How old are bacterial pathogens? Proc. Biol. Sci. 283, 20160990 (2016).
Kogan, A. et al. Common MEFV mutations among Jewish ethnic groups in Israel: high frequency of carrier and phenotype III states and absence of a perceptible biological advantage for the carrier state. Am. J. Med. Genet. 102, 272–276 (2001).
Ben-Chetrit, E. & Touitou, I. Familial Mediterranean fever in the world. Arthritis Rheum. 61, 1447–1453 (2009).
Ben-Chetrit, E. & Levy, M. Familial Mediterranean fever. Lancet 351, 659–664 (1998).
Zhao, Y. & Shao, F. Diverse mechanisms for inflammasome sensing of cytosolic bacteria and bacterial virulence. Curr. Opin. Microbiol. 29, 37–42 (2016).
Remmers, E. F. et al. Genome-wide association study identifies variants in the MHC class I, IL10, and IL23R-IL12RB2 regions associated with Behçet’s disease. Nat. Genet. 42, 698–702 (2010).
Barrett, J. C., Fry, B., Maller, J. & Daly, M. J. Haploview: analysis and visualization of LD and haplotype maps. Bioinformatics 21, 263–265 (2005).
Loh, P. R., Palamara, P. F. & Price, A. L. Fast and accurate long-range phasing in a UK Biobank cohort. Nat. Genet. 48, 811–816 (2016).
Sabeti, P. C. et al. Detecting recent positive selection in the human genome from haplotype structure. Nature 419, 832–837 (2002).
Szpiech, Z. A. & Hernandez, R. D. selscan: an efficient multithreaded program to perform EHH-based scans for positive selection. Mol. Biol. Evol. 31, 2824–2827 (2014).
Mallick, S. et al. The Simons Genome Diversity Project: 300 genomes from 142 diverse populations. Nature 538, 201–206 (2016).
The 1000 Genomes Project Consortium. A map of human genome variation from population-scale sequencing. Nature 467, 1061–1073 (2010).
Chae, J. J. et al. Targeted disruption of pyrin, the FMF protein, causes heightened sensitivity to endotoxin and a defect in macrophage apoptosis. Mol. Cell 11, 591–604 (2003).
Chae, J. J. et al. The B30.2 domain of pyrin, the familial Mediterranean fever protein, interacts directly with caspase-1 to modulate IL-1β production. Proc. Natl Acad. Sci. USA 103, 9982–9987 (2006).
Acknowledgements
We thank the patients enrolled in our clinical protocols for providing research specimens, A. Jones, T. Romeo, L. Poe and R. Laird for help with caring for patients and A. Schaffer for thoughtful discussions. This work was supported by the Intramural Research Programs of the National Human Genome Research Institute, the National Institute of Allergy and Infectious Diseases, the National Institute of Arthritis and Musculoskeletal and Skin Diseases and the Center for Research on Genomics and Global Health. This research was also supported by the National Institute of Allergy and Infectious Diseases award R01AI099222 (to J.B.B.) and utilized the computational resources of the NIH HPC Biowulf cluster (http://hpc.nih.gov).
Author information
Authors and Affiliations
Contributions
J.J.C. conceived the project. Y.H.P., E.F.R., J.B.B., D.L.K. D.S. and J.J.C. designed the study. Y.H.P., E.F.R., W.L., L.K.C., M.I.I., N.A.L., Y.Z.A.-U., B.B.-P., D.S. and J.J.C. performed experiments. Y.H.P., E.F.R., W.L., Z.S., I.A., C.N.R., H.C., J.B.B., D.L.K., D.S. and J.J.C. analyzed the data. Y.H.P., E.F.R., I.A., C.N.R., J.B.B., D.L.K., D.S. and J.J.C. wrote the manuscript. A.K.O., D.L.S., K.S.B., P.H., M.N., E.S., I.A., A.G. and S.O. provided the clinical data and specimens for healthy controls, carriers and patients. B.B.-P. and S.Z. provided laboratory resources and assistance at the Hacettepe University Faculty of Medicine.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Zoltan Fehervari was the primary editor on this article and managed its editorial process and peer review in collaboration with the rest of the editorial team.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
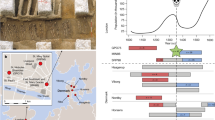
Extended Data Fig. 1 Haplotypes from 2,313 Turkish individuals provide evidence of evolutionarily recent positive selection and estimates of selection intensity and duration of selection on FMF-associated pyrin mutations.
a-c, Histograms of unstandardized nSL statistics of GWAS variants37 with similar frequency and local recombination rate as each FMF mutation. a, 1,402 markers similar to MEFV_p.V726A; b, 2,380 markers similar to MEFV_p.M694V; and c, 5,442 markers similar to MEFV_p.E148Q. The most extreme negative nSL values have the strongest evidence of recent positive selection. d-f, Log-likelihood plots and selection co-efficient estimates with 95% confidence intervals and mutation age estimates with 95% confidence intervals of three MEFV mutations, determined with a hidden Markov model method from the multi-locus haplotype structure of 4,626 Turkish chromosomes. d, MEFV_p.V726A; e, MEFV_p.M694V; and f, MEFV_p.E148Q. g-i, Histograms of iHH values for the ancestral alleles of markers with similar allele frequency and local recombination rate as each MEFV mutation, demonstrating that the ancestral alleles have not been under positive selection, thereby providing evidence that the mutations have not been under balancing selection. g, 1,570 markers similar to MEFV_ p.V726, h, 2,677 markers similar to MEFV_p.M694, and i, 6,166 markers similar to MEFV_p.E148.
Extended Data Fig. 2 Geographically diverse FMF mutation carriers exhibit extended conserved haplotypes.
a, MEFV_p.M694V conserved haplotypes (yellow). The length of the MEFV_p.M694V haplotype shared by all the diverse carrier chromosomes = 48 kb and shared by greater than 50% of the diverse carrier chromosomes = 307 kb. b, MEFV_p.V726A conserved haplotypes (pink). The length of the MEFV_p.V726A haplotype shared by all the diverse carrier chromosomes = 77 kb and shared by greater than 50% of the diverse carrier chromosomes = 262 kb. The fully conserved shared haplotypes in the diverse mutation carriers are identical to the respective haplotypes in the Turkish population.
Extended Data Fig. 3 Y. pestis YopM suppresses the pyrin inflammasome.
a,b, IL-1β measurements of culture supernatants of Mefv+/+ BMDMs primed with LPS, treated with or without C3 toxin, and infected with indicated MOI of WT (a) or ∆yopM (b) Y. pestis strains. Results are presented as mean ± s.e.m., for n = 5 independent biological replicates. c, Immunoblot analysis of pyrin and pro-IL-1β in lysates of retroviral transduced U937 cells, expressing WT or indicated Ser to Ala mutant pyrin proteins. Data are representative of three independent experiments with similar results.
Extended Data Fig. 4 RSK-YopM mediated pyrin phosphorylation is independent of PKN.
In vitro kinase assay of purified Myc/His-tagged N-terminal human pyrin (amino acids 1–330) incubated with recombinant PKN1 and/or RSK1 (a) or PKN2 and/or RSK1 (b) in the presence of purified GST or GST-YopM, and analyzed by immunoblot with antibody specific for S242 phosphorylated pyrin (a) or phosphorylated serine (b). Data are representative of three independent experiments with similar results.
Extended Data Fig. 5 5 Human and murine RSK isoforms exhibit a restricted distribution of gene expression in leukocytes.
a,b, Relative gene expression profile of RSK isoforms in human PBMCs with or without Y. pestis infection (a) and mouse BMDMs with or without LPS priming (b), and assayed by RT-QPCR. Results are presented as mean ± s.e.m., for n = 5 independent biological replicates.
Extended Data Fig. 6 Knockdown of RSK1, RSK2 and RSK3 in THP-1 cells induces ASC oligomerization.
a, Immunoblot analysis of lysates of THP-1 cells transiently transfected with negative control siRNA with no substantial sequence similarity to human gene sequences (N.C.) or a mixture of siRNAs targeting RSK1, RSK2 and RSK3. b, ASC oligomerization analysis by immunoblot from THP-1 cells transfected with N.C. or a mixture of siRNAs targeting RSK1, RSK2 and RSK3, and infected with Y. pestis (MOI 30). Cell lysates and the disuccinimidyl suberate (DSS)-treated pellets were analyzed by immunoblot with anti-ASC antibody. Data are representative of three independent experiments with similar results.
Extended Data Fig. 7 YopM binds to multiple regions of N-terminal pyrin.
a, Schematic structure of human pyrin with various deletion fragments of N-terminal human pyrin. b,c, GST-pulldown assay of V5-tagged various N-terminal human pyrin fragments with purified GST-YopM. Data are representative of three independent experiments with similar results.
Supplementary information
Supplementary Information
Supplementary Tables 1–3.
Source data
Source Data Fig. 1
Statistical source data.
Source Data Fig. 2
Statistical source data.
Source Data Fig. 2
Unprocessed immunoblots.
Source Data Fig. 3
Statistical source data.
Source Data Fig. 3
Unprocessed immunoblots.
Source Data Fig. 4
Unprocessed immunoblots.
Source Data Fig. 5
Statistical source data.
Source Data Fig. 5
Unprocessed immunoblots.
Source Data Fig. 6
Statistical source data.
Source Data Fig. 7
Statistical source data.
Source Data Fig. 7
Unprocessed immunoblots.
Source Data Fig. 8
Statistical source data.
Source Data Extended Data Fig. 1
Statistical source data.
Source Data Extended Data Fig. 3
Statistical source data.
Source Data Extended Data Fig. 3
Unprocessed immunoblots.
Source Data Extended Data Fig. 4
Unprocessed immunoblots.
Source Data Extended Data Fig. 5
Statistical source data.
Source Data Extended Data Fig. 6
Unprocessed immunoblots.
Source Data Extended Data Fig. 7
Unprocessed immunoblots.
Rights and permissions
About this article
Cite this article
Park, Y.H., Remmers, E.F., Lee, W. et al. Ancient familial Mediterranean fever mutations in human pyrin and resistance to Yersinia pestis. Nat Immunol 21, 857–867 (2020). https://doi.org/10.1038/s41590-020-0705-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41590-020-0705-6
This article is cited by
-
Genetically transitional disease: conceptual understanding and applicability to rheumatic disease
Nature Reviews Rheumatology (2024)
-
Serum endocan, asymmetric dimethylarginine and lipid profile in children with familial Mediterranean fever
Pediatric Research (2024)
-
Mutations in the B30.2 and the central helical scaffold domains of pyrin differentially affect inflammasome activation
Cell Death & Disease (2023)
-
Pathogenic Interleukin-10 Receptor Alpha Variants in Humans — Balancing Natural Selection and Clinical Implications
Journal of Clinical Immunology (2023)
-
Cardiovascular manifestations of monogenic periodic fever syndromes
Clinical Rheumatology (2023)