Abstract
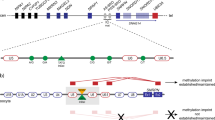
Imprinting on human chromosome 15 is regulated by an imprinting centre, which has been mapped to a 100–kb region including exon 1 of SNRPN. From this region we have identified novel transcripts, which represent alternative transcripts of the SNRPN gene. The novel exons lack protein coding potential and are expressed from the paternal chromosome only. We have also identified intragenic deletions and a point mutation in patients who have Angelman or Prader–Willi syndrome due to a parental imprint switch failure. This suggests that imprint switching on human chromosome 15 may involve alternative SNRPN transcripts.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout
Similar content being viewed by others
References
Glenn, C.C., Porter, K.A., Jong, M.T.C., Nicholls, R.D. & Driscoll, D.J. Functional imprinting and epigenetic modification of the human SNRPN gene. Hum. Mol. Genet. 2, 2001–2005 (1993).
Nakao, M. et al. Imprinting analysis of three genes in the Prader-Willi/Angelman region: SNRPN, E6-associated protein, and PAR-2 (D15S225E). Hum. Mol. Genet. 3, 309–315 (1994).
Reed, M. & Leff, S. Maternal imprinting of human SNRPN, a gene deleted in Prader-Willi syndrome. Nature Genet. 6, 163–167 (1994).
Glenn, C.C. et al. Gene structure, DNA methylation and imprinted expression of the human SNRPN gene. Am. J. Hum. Genet. 58, 335–346 (1996).
Özcelik, T. et al. Small nuclear ribonucleoprotein polypeptide N (SNRPN), an expressed gene in the Prader-Willi syndrome critical region. Nature Genet. 2, 265–269 (1992).
Sutcliffe, J.S. et al. Deletions of a differentially methylated CpG island at the SNRPN gene define a putative imprinting control region. Nature Genet. 8, 52–58 (1994).
Wevrick, R., Kerns, J.A. & Francke, U. Identification of a novel paternally expressed gene in the Prader-Willi syndrome region. Hum. Mol. Genet. 3, 1877–1882 (1994).
Razin, A. & Cedar, H. DNA methylation and genomic imprinting. Cell 77, 473–476 (1994).
Driscoll, D.J. et al. A DNA methylation imprint, determined by the sex of the parent, distinguishes the Angelman and Prader-Willi syndromes. Genomics 13, 917–924 (1992).
Dittrich, B. et al. Molecular diagnosis of the Prader-Willi and Angelman syndromes by detection of parent-of-origin specific DNA methylation in 15q11–13. Hum. Genet. 90, 313–315 (1992).
Dittrich, B., Suiting, K., Groβ, S. & Horsthemke, B. Characterization of a methylation imprint in the Prader-Willi syndrome region. Hum. Mol. Genet. 2, 1995–1999 (1993).
Suiting, K. et al. Inherited microdeletions in the Angelman and Prader-Willi syndromes define an imprinting centre on human chromosome 15. Nature Genet. 9, 395–400 (1995).
Glenn, C.C. et al. Modification of 15q11–q13 DNA methylation imprints in unique Angelman and Prader-Willi patients. Hum. Mol. Genet. 2, 1377–1382 (1993).
Buiting, K. et al. Detection of aberrant DNA methylation in unique Prader-Willi syndrome patients and ist diagnostic implications. Hum. Mol. Genet. 3, 893–895 (1994).
Ledbetter, D. et al. Deletions of chromosome 15 as a cause of the Prader-Willi syndrome. New Engl. J. Med. 304, 325–329 (1981).
Knoll, J.H.M. et al. Angelman and Prader-Willi syndrome share a common chromosome 15 deletion but differ in parental origin of the deletion. Am. J. Med. Genet. 32, 285–290 (1989).
Nicholls, R.D., Knoll, J.H.M., Butler, M.G., Karam, S. & Lalande, M. Genetic imprinting suggested by maternal heterodisomy in non-deletion Prader-Willi syndrome. Nature 342, 281–285 (1989).
Reis, A. et al. Imprinting mutations suggested by abnormal DNA methylation patterns in familial Angelman and Prader-Willi syndromes. Am. J. Hum. Genet. 54, 741–747 (1994).
Kaplan, L.C. et al. Clinical heterogeneity associated with deletions in the long arm of chromosome 15: report of 3 new cases and their possible significance. Am. J. Med. Genet. 28, 45–53 (1987).
Magenis, R.E., Brown, M.G., Lacy, D.A., Budden, S. & LaFranchi, S. Is Angelman syndrome an alternate result of del(15)(q11 q13)? Am. J. Med. Genet. 28, 829–838 (1987).
Pembrey, M. et al. The association of AngelmanOs syndrome with deletions within 15q11–13. J. Med. Genet. 26, 73–77 (1989).
Malcolm, S. et al. Uniparental paternal disomy in Angelman's syndrome. Lancet 337, 694–697 (1991).
Clayton-Smith, J. et al. Further evidence for dominant inheritance at the chromosome 15q11-13 locus in familial Angelman syndrome. Am. J. Hum. Genet. 44, 256–260 (1992).
Wagstaff, J. et al. Maternal but not paternal transmission of 15q11–q13-linked nondeletion Angelman syndrome leads to phenotypic expression. Nature Genet. 1, 291–294 (1992).
Saitoh, S. et al. Minimal definition of the imprinting centre and fixation of a chromosome 15q11–13 epigenotype by imprinting mutations. Proc. Natl. Acad. Sci USA 93, 7811–7815
Saitoh, S. et al. Familial Angelman syndrome caused by imprinted submicroscopic deletion encompassing GABAA receptor b3-subunit gene. Lancet 339, 366–367 (1992).
Wagstaff, J., Shugart, Y.Y. & Lalande, M. Linkage analysis in familial Angelman syndrome. Am. J. Hum. Genet. 53, 105–112 (1993).
Horsthemke, B., Dittrich, B. & Buiting, K. Parent-of-origin specific DNA methylation and imprinting mutations on human chromosome 15. in Parental imprinting: causes and consequences (eds Ohlsson, R., Hall, K. & Ritzen, M.) 295–308 (Cambridge University Press, 1995).
Korn, B. et al. A strategy for the selection of transcribed sequences in theXq28 region. Hum. Mol. Genet. 1, 235–242 (1992).
Brockdorff, N. et al. The product of the mouse Xist gene is a 15-kb inactive X-specific transcript containing no conserved ORF and located in the nucleus. Cell 71, 515–526 (1992).
Orlando, V. & Paro, R. Chromatin multiprotein complexes involved in the maintenance of transcription patterns. Curr. Opin. Genet. Dev. 5, 174–179 (1995).
Reik, W. et al. Imprinting mutations in the Beckwith-Wiedemann syndrome suggested by an altered imprinting pattern in the IGF2-H19 domain. Hum. Mol. Genet. 4, 2379–2385 (1995).
Chaillet, J.R., Knoll, J.H., Horsthemke, B. & Lalande, M. The syntenic relationship between the critical deletion region for the Prader-Willi/Angelman syndromes and proximal mouse chromosome 7. Genomics 11, 773–776 (1991).
Nicholls, R.D. et al. Evaluation of potential models for imprinted and nonimprinted components of human chromosome 15q11–q13 syndromes by fine-structure homology mapping in the mouse. Proc. Natl. Acad. Sci. USA 90, 2050–2054 (1993).
Chan, C.T.J. et al. Molecular mechanisms in Angelman syndrome: a survey of 93 patients. J. Med. Genet. 30, 895–902 (1993).
Mutirangura, A. et al. A complete YAC contig of the Prader-Willi/Angelman chromosome region (15q11–q13) and refined localization of the SNRPN gene. Genomics 18, 546–552 (1993).
Horsthemke, B., Greger, V., Barnert, H.J., HÖpping, W. & Passarge, E. Detection of submicroscopic deletions and a DNA polymorphism at the retinoblastoma locus. Hum. Genet. 76, 257–261 (1987).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Dittrich, B., Buiting, K., Korn, B. et al. Imprint switching on human chromosome 15 may involve alternative transcripts of the SNRPN gene. Nat Genet 14, 163–170 (1996). https://doi.org/10.1038/ng1096-163
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/ng1096-163
This article is cited by
-
The inheritance of epigenetic defects
medizinische genetik (2017)
-
Pharmacological therapies for Angelman syndrome
Wiener Medizinische Wochenschrift (2017)
-
Differential regulation of non-protein coding RNAs from Prader-Willi Syndrome locus
Scientific Reports (2014)
-
Identification and characterization of alternative exon usage linked glioblastoma multiforme survival
BMC Medical Genomics (2012)