Abstract
Elevated basal serum tryptase levels are present in 4–6% of the general population, but the cause and relevance of such increases are unknown1,2. Previously, we described subjects with dominantly inherited elevated basal serum tryptase levels associated with multisystem complaints including cutaneous flushing and pruritus, dysautonomia, functional gastrointestinal symptoms, chronic pain, and connective tissue abnormalities, including joint hypermobility. Here we report the identification of germline duplications and triplications in the TPSAB1 gene encoding α-tryptase that segregate with inherited increases in basal serum tryptase levels in 35 families presenting with associated multisystem complaints. Individuals harboring alleles encoding three copies of α-tryptase had higher basal serum levels of tryptase and were more symptomatic than those with alleles encoding two copies, suggesting a gene-dose effect. Further, we found in two additional cohorts (172 individuals) that elevated basal serum tryptase levels were exclusively associated with duplication of α-tryptase–encoding sequence in TPSAB1, and affected individuals reported symptom complexes seen in our initial familial cohort. Thus, our findings link duplications in TPSAB1 with irritable bowel syndrome, cutaneous complaints, connective tissue abnormalities, and dysautonomia.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
Accession codes
References
Fellinger, C. et al. Clinical characteristics and risk profile of patients with elevated baseline serum tryptase. Allergol. Immunopathol. (Madr.) 42, 544–552 (2014).
Gonzalez-Quintela, A. et al. Factors influencing serum total tryptase concentrations in a general adult population. Clin. Chem. Lab. Med. 48, 701–706 (2010).
Henningsen, P., Zimmermann, T. & Sattel, H. Medically unexplained physical symptoms, anxiety, and depression: a meta-analytic review. Psychosom. Med. 65, 528–533 (2003).
Cardet, J.C., Castells, M.C. & Hamilton, M.J. Immunology and clinical manifestations of non-clonal mast cell activation syndrome. Curr. Allergy Asthma Rep. 13, 10–18 (2013).
Li, H. et al. Autoimmune basis for postural tachycardia syndrome. J. Am. Heart Assoc. 3, e000755 (2014).
Zarate, N. et al. Unexplained gastrointestinal symptoms and joint hypermobility: is connective tissue the missing link? Neurogastroenterol. Motil. 22, 252–e78 (2010).
Lyons, J.J. et al. Mendelian inheritance of elevated serum tryptase associated with atopy and connective tissue abnormalities. J. Allergy Clin. Immunol. 133, 1471–1474 (2014).
Ross, J. & Grahame, R. Joint hypermobility syndrome. Br. Med. J. 342, c7167 (2011).
Buskila, D. & Sarzi-Puttini, P. Biology and therapy of fibromyalgia. Genetic aspects of fibromyalgia syndrome. Arthritis Res. Ther. 8, 218 (2006).
Castori, M. Ehlers–Danlos syndrome, hypermobility type: an underdiagnosed hereditary connective tissue disorder with mucocutaneous, articular, and systemic manifestations. ISRN Dermatol. 2012, 751768 (2012).
Sabato, V. et al. Familial hypertryptasemia with associated mast cell activation syndrome. J. Allergy Clin. Immunol. 134, 1448–1450.e3 (2014).
Saito, Y.A., Schoenfeld, P. & Locke, G.R., III. The epidemiology of irritable bowel syndrome in North America: a systematic review. Am. J. Gastroenterol. 97, 1910–1915 (2002).
El-Serag, H.B., Sweet, S., Winchester, C.C. & Dent, J. Update on the epidemiology of gastro-oesophageal reflux disease: a systematic review. Gut 63, 871–880 (2014).
Remvig, L., Jensen, D.V. & Ward, R.C. Epidemiology of general joint hypermobility and basis for the proposed criteria for benign joint hypermobility syndrome: review of the literature. J. Rheumatol. 34, 804–809 (2007).
Ruëff, F. et al. Predictors of severe systemic anaphylactic reactions in patients with Hymenoptera venom allergy: importance of baseline serum tryptase—a study of the European Academy of Allergology and Clinical Immunology Interest Group on Insect Venom Hypersensitivity. J. Allergy Clin. Immunol. 124, 1047–1054 (2009).
Sturm, G.J. et al. Sensitization to Hymenoptera venoms is common, but systemic sting reactions are rare. J. Allergy Clin. Immunol. 133, 1635–43 e1 (2014).
Savova, V. et al. Genes with monoallelic expression contribute disproportionately to genetic diversity in humans. Nat. Genet. 48, 231–237 (2016).
Bailey, J.A. et al. Recent segmental duplications in the human genome. Science 297, 1003–1007 (2002).
Chiang, P.W. et al. Somatic and germ-line mosaicism in Rubinstein–Taybi syndrome. Am. J. Med. Genet. A. 149A, 1463–1467 (2009).
Trivedi, N.N., Tong, Q., Raman, K., Bhagwandin, V.J. & Caughey, G.H. Mast cell α and β tryptases changed rapidly during primate speciation and evolved from γ-like transmembrane peptidases in ancestral vertebrates. J. Immunol. 179, 6072–6079 (2007).
Drossman, D.A. & Dumitrascu, D.L. Rome III: new standard for functional gastrointestinal disorders. J. Gastrointestin. Liver Dis. 15, 237–241 (2006).
Sletten, D.M., Suarez, G.A., Low, P.A., Mandrekar, J. & Singer, W. COMPASS 31: a refined and abbreviated Composite Autonomic Symptom Score. Mayo Clin. Proc. 87, 1196–1201 (2012).
Biesecker, L.G. et al. The ClinSeq Project: piloting large-scale genome sequencing for research in genomic medicine. Genome Res. 19, 1665–1674 (2009).
Alfter, K. et al. New aspects of liver abnormalities as part of the systemic mast cell activation syndrome. Liver Int. 29, 181–186 (2009).
Zhang, Y. et al. Autosomal recessive phosphoglucomutase 3 (PGM3) mutations link glycosylation defects to atopy, immune deficiency, autoimmunity, and neurocognitive impairment. J. Allergy Clin. Immunol. 133, 1400–1409, 1409.e1–1409.e5 (2014).
Brennan, C.W. et al. The somatic genomic landscape of glioblastoma. Cell 155, 462–477 (2013).
Johnston, J.J. et al. Individualized iterative phenotyping for genome-wide analysis of loss-of-function mutations. Am. J. Hum. Genet. 96, 913–925 (2015).
Abecasis, G.R., Cherny, S.S., Cookson, W.O. & Cardon, L.R. Merlin—rapid analysis of dense genetic maps using sparse gene flow trees. Nat. Genet. 30, 97–101 (2002).
Kristensen, T., Vestergaard, H., Bindslev-Jensen, C., Møller, M.B. & Broesby-Olsen, S. Sensitive KIT D816V mutation analysis of blood as a diagnostic test in mastocytosis. Am. J. Hematol. 89, 493–498 (2014).
Ferrer, M. et al. Serum total tryptase levels are increased in patients with active chronic urticaria. Clin. Exp. Allergy 40, 1760–1766 (2010).
Schwartz, L.B. Diagnostic value of tryptase in anaphylaxis and mastocytosis. Immunol. Allergy Clin. North Am. 26, 451–463 (2006).
Le, Q.T., Lotfi-Emran, S., Min, H.K. & Schwartz, L.B. A simple, sensitive and safe method to determine the human α/β-tryptase genotype. PLoS One 9, e114944 (2014).
Miller, J.S., Moxley, G. & Schwartz, L.B. Cloning and characterization of a second complementary DNA for human tryptase. J. Clin. Invest. 86, 864–870 (1990).
Trivedi, N.N., Tamraz, B., Chu, C., Kwok, P.Y. & Caughey, G.H. Human subjects are protected from mast cell tryptase deficiency despite frequent inheritance of loss-of-function mutations. J. Allergy Clin. Immunol. 124, 1099–105.e1, 4 (2009).
Kirshenbaum, A.S. et al. Demonstration that human mast cells arise from a progenitor cell population that is CD34+, c-kit+, and expresses aminopeptidase N (CD13). Blood 94, 2333–2342 (1999).
Schwartz, L.B., Lewis, R.A., Seldin, D. & Austen, K.F. Acid hydrolases and tryptase from secretory granules of dispersed human lung mast cells. J. Immunol. 126, 1290–1294 (1981).
Tkaczyk, C., Metcalfe, D.D. & Gilfillan, A.M. Determination of protein phosphorylation in FcɛRI-activated human mast cells by immunoblot analysis requires protein extraction under denaturing conditions. J. Immunol. Methods 268, 239–243 (2002).
Acknowledgements
We thank the patients, their families, and the numerous healthy volunteers who contributed to this research, as well as the clinical staff of the LAD, NIMH, NIDDK, and NHGRI for their efforts, in particular the Gastrointestinal and Psychiatry Consultation Liaison Services physicians, especially T. Alqassem, who participated in the care of these patients. We thank all the referring care providers, in particular A. Maitland and M. Carter for each referring several families for evaluation. We also acknowledge the collaborative spirit and efforts of the NIAMS and NIAID clinical genomics programs, specifically the investigators (M.J. Lenardo, H.S. Su, and R.T. Goldbach-Mansky) who shared clinical genomics data and study samples. We also thank M.J. Lenardo, W. Gahl, C. Akin, and J.-L. Casanova for their review of the manuscript. Lastly, we thank D. Abdulazeez of VCU for performing the tryptase immunoassays. This study was supported in part by the Division of Intramural Research of the National Institute of Allergy and Infectious Diseases, NIH. The involvement of N.J. was funded by NCI contract HHSN261200800001E. Funding was also provided in part by ARTrust/The Mastocytosis Society Research Award in Mastocytosis and/or Mast Cell Activation Syndrome (J.J.L.) and by NIH HL024136 (G.H.C.). L.G.B., C.H., and K.L.L. were supported by the Intramural Research Program of the NHGRI.
Author information
Authors and Affiliations
Contributions
J.J.L. and J.D.M. designed the study. J.J.L., J.D.M., C.N., T.D., N.J., H.M., T.M.W., K.D.S., D.D.M., S.C.G., P.D.A., and M.E.R. all recruited subjects to the study. J.J.L., J.D.M., T.H., C.N., N.J., T.D., N.H., M.P., S.C.G., R.J.C., S.A., S.T., T.M.W., and A.J. collected and/or analyzed clinical data. X.Y., J.D.H., C.H., Y.Z., A.J.O., J.J.M., L.G.B., and J.D.M. performed and supported genomic sequencing. X.Y., J.D.H., C.H., Y.Z., and A.J.O. performed the bioinformatic analyses. J.J.L., N.N.T., G.H.C., and L.B.S. designed and J.J.L. performed the ddPCR assay. J.J.L., Q.T.L., Y.B., M.Z., Y.L., M.P.O'C., R.J.H., L.B.S., and J.D.M. designed and performed the functional studies. K.L.L., C.H., and L.G.B. facilitated all ClinSeq-related studies. J.J.L. and J.D.M. prepared the draft manuscript. All authors contributed to discussion of the results and to manuscript preparation.
Corresponding author
Ethics declarations
Competing interests
The NIH authors declare no conflicts of interest. L.B.S. receives royalties from VCU that are collected from Thermo Fisher for the tryptase UniCAP assay and receives consulting fees from Genentech, Inc. L.G.B. is an uncompensated advisor to Illumina and receives royalties from Genentech, Inc., and Amgen, and honoraria from Wiley-Blackwell.
Integrated supplementary information
Supplementary Figure 1 Study design for identification of hereditary α-tryptasemia, characterization of associated clinical features, and confirmation of the genetic and clinical features in additional populations.
(a) Schematic for evaluation of the referral α-tryptasemia cohort. Families were referred for symptomatic elevation of basal serum tryptase levels without mastocytosis or for familial connective tissue abnormalities in the context of atopy and/or symptoms often associated with mast cell mediators. (b) Schematic for evaluation of the NIAID/NIAMS (left) and ClinSeq (right) cohorts. To enrich for individuals with elevated tryptase levels, exome data were reviewed in the 951 individuals enrolled in ClinSeq. Limited coverage of the 16p13.3 locus permitted the selection of 33 individuals with single-nucleotide variants (SNVs) in genes adjacent to TPSAB1 (TPSG1 (rs113856625[G>A]) and CACNA1H (rs58124832[G>A]) out of 513 with ≥10× coverage at these loci. These SNVs were observed to segregate in 8 of 12 families sequenced with hereditary α-tryptasemia syndrome. Neither SNV has been reported to cause disease; the SNV in CACNA1H in combination with another variant has been reported in association with autism spectrum disorders, which were not seen in this cohort (J. Biol. Chem. 281, 22085–22091, 2006). Given that the minor allele frequency (MAF) of these SNVs in Caucasians is approximately 0.06, enrichment was estimated to be between two- and fourfold. An additional 92 patients without these SNVs were selected at random. A cutoff basal serum tryptase level of ≥8 ng/ml was established for further genetic testing based on the range of tryptase levels in the 96 individuals identified with hereditary α-tryptasemia syndrome and the additional 8 individuals identified with hereditary α-tryptasemia in the first and second cohorts, respectively (8–39.5 ng/ml). Of the 25 individuals with basal serum tryptase concentration ≥8 ng/ml, sufficient exome sequence was captured within the TPSAB1 locus itself to perform bioinformatic genotyping in 16 individuals; the remaining 9 samples had to be excluded. A total of nine individuals were identified with TPSAB1 duplication of α-tryptase–encoding sequences, seven of whom carried the CACNA1H and TPSG1 variants and two of whom did not. Absence of TPSAB1 duplication was confirmed by bioinformatic analysis in all 65 individuals with basal serum tryptase concentration <8 ng/ml for whom capture of TPSAB1 sequence permitted analysis.
Supplementary Figure 2 Increased copy number of α-tryptase–encoding sequences in TPSAB1 is inherited in families with α-tryptasemia and is associated with higher dysautonomia scores.
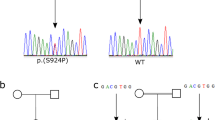
(a) Distribution of total autonomic dysfunction scores from affected individuals (α-tryptasemia, n = 70) and unaffected family members (control, n = 31) with inherited TPSAB1 duplications and triplications, obtained using the standardized COMPASS 31 questionnaire. Data are shown as medians ± interquartile range, Mann–Whitney test. (b) Five sample pedigrees showing dominant inheritance of multiple α-tryptase–encoding TPSAB1 gene copies based on α-tryptase/β-tryptase ratios, which segregate with elevated basal serum tryptase levels (filled symbols). Individuals with normal basal serum tryptase levels are represented with open symbols. (c) α-tryptase/β-tryptase ratios obtained from individuals with basal serum tryptase concentration >11.4 ng/ml (affected) and those with normal basal serum tryptase concentrations (unaffected) from 15 families with dominantly inherited elevation of basal serum tryptase levels.
Supplementary Figure 3 Consensus α-tryptase–encoding sequence and in silico alignment.
(a) Consensus α-tryptase–encoding sequence derived in silico. (b) Alignment of the sequences encoding α-, βI/II-, βIII-, and δ-tryptase with primer (turquoise), probe (yellow), and restriction site (blue) sequences highlighted.
Supplementary Figure 4 Schematic of the screening bioinformatics algorithm used to estimate copy number of α-tryptase–encoding sequences in TPSAB1.
Genomic DNA sequence reads that mapped initially to the general tryptase locus (chr. 16: 1,250,000–1,350,000) were remapped to the α-tryptase-encoding consensus sequence derived in silico. Using a computer algorithm, unique sequence ‘clusters’ with complete internal sequence homology were identified. These clusters were then assigned to one of the unique gene sequences encoding α-, βI/II-, βIII-, or δ-tryptase. On the basis of the number of unique sequences, the number of clusters mapping back to each specific gene, and the number of reads (read #) covering that sequence, an estimated copy number (Est. copy) could be obtained for each gene sequence. Using these estimates, the α-tryptase/β-tryptase genotype encoded at the TPSAB1 and TPSB2 loci could be predicted.
Supplementary Figure 5 Digital droplet PCR assay of α- and β-tryptase.
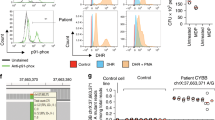
(a) One-dimensional plots of ddPCR data obtained from the genomic DNA of two individuals. Left panel, two individuals, one with an allele encoding α-tryptase at TPSAB1 and positive (blue) droplets (right) and one with only β-tryptase-encoding alleles at TPSAB1 (left). Right panel, droplets containing reference gene AP3B1 for both individuals (green). (b) Concentration of α-tryptase–encoding (blue) and AP3B1 (green) genomic DNA in copies per microliter (left) and corresponding copy number calls for α-tryptase–encoding sequences (right); the genotypes of the four samples are indicated at the bottom. When duplication (αα) or triplication (ααα) of a TPSAB1 gene encoding α-tryptase is present on a single allele (within 50 kb), a shift (Δα) in the concentration of α-tryptase–encoding sequence relative to the reference is seen with restriction digestion by BamHI, resulting in an increase in calculated copy number (Δcopy), whereas when two different alleles encode α-tryptase, no shift is seen (arrows). (c) The ratio of alleles encoding β-tryptase (green) relative to those encoding α-tryptase (blue) (left) also increases (Δβ) following BamHI digestion if two copies of a β-tryptase–encoding sequence are present on a single allele, resulting in a change (arrows) in the α/β ratio (Δratio), thereby allowing for determination of complete tryptase allele genotypes at TPSAB1 and TPSB2 (bottom right). (d) Histograms of raw copy number calls for α-tryptase–encoding sequences in TPSAB1 from individuals with hereditary α-tryptasemia (left) and unaffected family members (right), with and without restriction digestion. Individuals with duplications and triplications of α-tryptase–encoding TPSAB1 sequences in cis (on the same allele) initially had an artificially low copy number call owing to droplets containing multiple α-tryptase–encoding sequences that did not independently sort (left). Following brief restriction digestion, a shift toward increased copy number was seen in these individuals, while individuals with two copies of α-tryptase–encoding sequences in trans (on separate alleles) demonstrated no change in copy number as detected by ddPCR (right).
Supplementary Figure 6 Cultured mast cells from individuals with α-tryptasemia express more tryptase transcript, while intracellular protein levels and degranulation activity appear to be normal.
(a) Total intracellular tryptase protein expression in mast cells cultured from the peripheral CD34+ cells of individuals with duplication or triplication of α-tryptase–encoding sequences in TPSAB1 (α-tryptasemia, n = 4) and paired cultures (control, n = 4) were measured by immunoblot (left; normalized to β-actin) and by flow cytometry following intracellular staining (right) (α-tryptasemia, n = 5, control, n = 6). Data are combined from five independent culture experiments and are shown as means ± s.e.m., Wilcoxon matched-pairs test. (b) Mast cell degranulation in response to increasing antigen (Ag) concentration was measured in cultured mast cells by β-hexosaminidase (β-hex) release in the presence or absence of human stem cell factor (hSCF). Data are from three independent culture experiments (α-tryptasemia, n = 4; control, n = 5) and are shown as means ± s.d. (c) Total TPSAB1 and TPSB2 transcripts (total tryptase) were measured in cultured mast cells (five independent paired cultures) and in total PBMCs (n = 10 versus 10) from individuals with inherited α-tryptase copy number increases (a -tryptasemia) or paired individuals without extra α-tryptase copies (control) by real-time PCR; data are shown as means ± s.e.m., Wilcoxon matched-pairs test.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–6, Supplementary Tables 1–4 and Supplementary Note. (PDF 1699 kb)
Rights and permissions
About this article
Cite this article
Lyons, J., Yu, X., Hughes, J. et al. Elevated basal serum tryptase identifies a multisystem disorder associated with increased TPSAB1 copy number. Nat Genet 48, 1564–1569 (2016). https://doi.org/10.1038/ng.3696
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.3696
This article is cited by
-
Clinically accessible amplitude-based multiplex ddPCR assay for tryptase genotyping
Scientific Reports (2024)
-
Mastozytose – eine häufig unerkannte Erkrankung
Die Dermatologie (2024)
-
Mast Cell Activation Syndrome and Gut Dysfunction: Diagnosis and Management
Current Gastroenterology Reports (2024)
-
Predictors of Clonality and Underlying Mastocytosis in Mast Cell Activation Syndromes
Current Allergy and Asthma Reports (2024)
-
Alpha-Tryptase as a Risk-Modifying Factor for Mast Cell–Mediated Reactions
Current Allergy and Asthma Reports (2024)